Short Communication
COMPLETE GENOME SEQUENCE OF ESCHERICHIA COLI E706 ISOLATED FROM A CHICKEN FARM
S. Zhao, G. R. Mustafa, S. Yang, W. Deng, X. Yu and L. Zou*
Department of Applied Microbiology, College of Resources, Sichuan Agricultural University, 611130, Chengdu, Sichuan, China
*Corresponding author’s email: zoulikou@sicau.edu.cn
ABSTRACT
Escherichia coli is normal gut commensal for animals as well as humans, meanwhile it can be present in the environment. Basing on whole genome sequencing (WGS), we characterized the E. coli strain E706, which harbors a series of virulence and resistance genes, CRISPR candidate elements, prophage related sequences, as well as genomic islands. The plasmid pLKJULY7989 in E706 is a chimera plasmid made up of a basic IncFIB plasmid backbone with class 1 integron and multi-resistance genes. pLKJULY7989 carries a class 1 integron with the existence of gene cassette dfrA12-aadA2 and class 1 integrase gene intl1, which is lacked in homologous plasmid pEHS30-1. The class 1 integron was detected only harboring a class 1 integrase gene, and gene cassette dfrA12-aadA2. The complete sequence we reported here will provide valuable information for a more understanding about the genomic structure and pathogenic characteristics of the genome of E. coli.
Keywords Escherichia coli, plasmid, whole genome sequencing, resistance gene.
https://doi.org/10.36899/JAPS.2021.3.0283
Published online November 09,2020
INTRODUCTION
Escherichia coli (E. coli) is normal gut commensal for animals as well as humans and animals, meanwhile it can be present the environment (Colello et al., 2018; Runcharoen et al., 2017). Several serotypes of E. coli can cause host disease (Zhu et al., 2017). As an emerging pathogen, E. coli often carries virulence factors that responsible for outbreaks of diarrhea (such as katP, etpD, efa/lifA and ent) (Zhang et al., 2018) and inflammatory bowel (such as fliC and fimH) (Palmela et al., 2018). The existence of multiple antibiotic resistance genes (ARGs) in E. coli could lead to the difficult treatment of the diseases (Flynn et al., 2015). Moreover, the presence of mobile elements, i.e., integrons, transposons and plasmids, in E. coli could cause serious threat by the dissemination of resistance genes to the environment (Blair, Webber, Baylay, Ogbolu, & Piddock, 2015). In addition to antibiotics, the heavy metal contamination triggers bacteria, including E. coli, Enterococcus faecium and Lactococcus garvieae, developed different strategies for their survival against antibiotics and metals. Similar to ARGs, the bacteria could develop heavy metal resistance genes (HMRGs) in response to heavy metals in their surroundings. The HMRGs can be situated on the chromosome, as well as plasmids of bacteria (Ture, Altinok, & Alp, 2018).
Plasmids belong to incompatibility F (IncF) group have frequently been linked with phenotype of multiple drug resistance (MDR), including plasmid-mediated quinolone resistance (PMQR) and extended spectrum lactamases (ESBLs) genes. In various studies, most of resistance determinants were carried by IncF plasmids, including floR, oqxAB, rmtB, and blaCTX-M (Lyimo et al., 2016; Matsumura et al., 2013). E. coli strains, that are capable to produce Extended spectrum β-lactamase (ESBL), have capacity of hydrolyzing various antibiotics, i.e., third-generation cephalosporin (Runcharoen et al., 2017).
Till now, whole genome sequencing (WGS) has mainly been used as a molecular research tool for pathogen detection (van El et al., 2013). With the development of WGS, antimicrobial and virulence resistance profile of bacteria can be predicted (Lindsey, Pouseele, Chen, Strockbine, & Carleton, 2016). There have been a lot of researches on complete sequence of different E. coli stains (Gilrane et al., 2017; Pan et al., 2016; Wong et al., 2016). Herein, we characterized the E. coli E706 using whole genome sequencing for a better understanding of its potential virulence and resistance genes.
MATERIALS AND METHODS
E706 strain isolation and identification: The E. coli E706 was isolated from air environment in a chicken farm located in southern west of China in 2017. The strain was isolated with Eosin Methylene Blue (EMB) agar. The DNA extraction of E. coli E706 was performed with Bacterial DNA Extraction Kit (Centrifugal column type DP2011) (BioTeke Inc., China). After confirmed by 16S rRNA sequencing, the strain was immediately frozen in glycerol at - 80 ℃ until use.
Whole genome sequencing of E706: The complete sequence of E. coli strain E706 was performed using third-generation sequencing based on PacBio platform (Illumina). Prediction of genome component comprised of the prophage, transposon, genomics islands, non-coding RNA, repetitive sequences, and coding gene were performed by NCBI Blast. The Blast search (E-value < 1e-5, minimal alignment length percentage > 40%) for gene prediction above was accomplished against Gene Ontology (GO; https://www.uniprot.org/help/gene_ontology), Kyoto Encyclopedia of Genes and Genomes (KEGG; ) (https://www.kegg.jp/), Clusters of Orthologous Groups (COG; http://www.ncbi.nlm.nih.gov/COG), Non-Redundant Protein Database databases (NR ; https://www.biostars.org), Transporter Classification Database (TCDB; http://www.tcdb.org/browse.php), and, Swiss-Prot (https://web.expasy.org/docs/swiss-prot) databases. Meanwhile, the Comprehensive Antibiotic Resistance Database (CARD; https://card.mcmaster.ca) was performed to analyze the drug resistance. In addition, the Virulence Factors of Pathogenic Bacteria (VFDB; http://www.mgc.ac.cn/VFs/) was used to perform the virulence analysis.
The type of plasmid was determined by PlasmidFinder 1.3 (Carattoli et al., 2014) (https://cge.cbs.dtu.dk/services/PlasmidFinder/). Genes were annotated by RAST (Brettin et al., 2015). Plasmid sequence comparisons were performed by using EasyFig 2.2.7 and the BLASTN (Sullivan, Petty, & Beatson, 2011).
RESULTS
Genome information and gene component of E706: Totally, 107,009 single reads of an average length of 11,634 bp were obtained in E706. After filtering of raw data, 2 contigs with an N50 of 2,981,102 bp were obtained. BLASTn search revealed that E. coli strain E706 is closely related to strain ECCWS199 (E. coli; accession CP032237.1; 90% coverage; 99% identity), isolated from chicken stool, and strain FORC_031 (E. coli; CP013190.1; 90% coverage; 99% identity) from human stool. The E. coli E706 has a single circular chromosome that was 4,696,154 bp in length, corresponding to 150-fold genome coverage, with GC content of 51.0% and encodes 83, 27 and 2 numbers of tRNA, rRNA and sRNA, respectively. 4,869 protein-coding genes were predicted. The genomic islands (GIs) were abundant in strain E706, which had 13 GIs with the average length of 14,702 bp. Most of coded proteins in GIs were described as hypothetical protein, but some GIs contained genes that are involved in acid stress, phage assemble and DNA replication. In addition, 2 Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) candidate elements and 13 prophage related sequences, ranging from 14 kb to 58 kb, were also detected. The general genomic features of E706 were visualized in Fig.1.
Virulence genes located on E706 chromosome: There are 269 putative virulence factors were predicted in E706. All the virulence genes belong to different gene families including fli, flg, fim, hem, che, ent, lpx, esp, csg, fep, ecp, flh, ccm, sta, stf, waa, alg, ycb, out, pil, clp, rfa, orf, nar, mbt, ibe, laf, muc, mgt, pan, mtr, rpo, pul, kdt, fle, feo, far, pho, gac, afr, omp, eps, gal, eae, mot, kat and eha as well as 68 individual genes (Online Resource 3). Compared with other E. coli isolates, E706 harbors a lot of common gene families such as pil, flh, fle, fli, bvgA and flg. These gene families are responsible for type IV pilus encoding, biofilm formation, Swimming motility and pathogenic adaptations. Meanwhile, E706 carries one virB gene which belongs to type IV secretion systems.
Resistance genes on E706 chromosome: A variety of ARGs were identified on the chromosome using BLAST together with the CARD database. Our strain E706 harbors 23 resistance genes, including emrE (aminoglycoside), emrD (aminoglycoside), mdfA (multidrug efflux pump), mdtF (doxorubicin/erythromycin), mdtG (deoxycholate/fosfomycin), mdtH (deoxycholate/fosfomycin), mdtK (enoxacin/norfloxacin), mdtL (chloramphenicol), mdtM (chloramphenicol/norfloxacin), mdtN, mdtO (puromycin/acriflavine), mdtP (puromycin/acriflavine), blaTEM-1 (penicillin), bacA (bacitracin), arnA (polymyxin), dfrA (trimethoprim), acrA (glycylcycline/acriflavin), acrB (glycylcycline/acriflavin), ksgA (kasugamycin), sul2 (sulfonamide), mphA (macrolide), tetC (tetracycline) and ant(2') (tobramycin/kanamycin/sisomicin/dibekacin/gentamicin). Besides, several heavy metal resistance genes, arsC (arsenic, As), copC (copper, Cu), copD (copper, Cu), corA (magnesium/nickel/cobalt, Mg/Ni/Co), mntH (manganese, Mg), nikB (nickel, Ni), nikC (nickel, Ni), nikD (nickel, Ni), nikE (nickel, Ni), corC (magnesium/cobalt, Mg/Co), terC (tellurium, Te) and czcD (Cobalt/zinc/cadmium, Co/Zn/Cd), were also included (Online Resource 1).
Other factors on E706 chromosome: Apart from 42 uncharacterized proteins, 3903 genes were annotated by COG databases. These genes encode carbohydrate metabolism and transportation, amino acid metabolism and transportation, energy conversion and production, and transcription respectively that accounted for a large proportion (each more than 12%), which indicated the better absorption capacity and response ability for amino acids and carbohydrates. In addition, high read coverage of E706 enabled methylation modifications to be annotated, and 168,296 m4C (4-methyl-cytosine) and 13,026 m6A (6-methyl-adenosine) residues were recognized.
Gene component in plasmid pLKJULY7989: The plasmid pLKJULY7989 was 151,880 bp in length with GC content of 49.5% and belonged to the IncFIB type. It comprised 169 open reading frames (orf). The plasmid pLKJULY7989 harbors floR (florfenicol/chloramphenicol resistance), strA (streptomycin resistance), strB (streptomycin resistance), tetR (tetracycline resistance), mphA (Erythromycin resistance), blaTEM-1 (ampicillin resistance), and sul2 (sulfonamide resistance) resistance gene. Moreover, this plasmid carries a heavy metal resistance gene (arsR) and a class 1 integron with gene cassette dfrA12- aadA2. Additionally, a diverse range of transposons, including IS1, S6, IS3, IS30, IS91, IS4, IS21, IS110, IS5, IS200 were identified. It was notable that the multiple resistance region flanked by IS6 and IS1comprised six resistance genes, including mph(A), tetR and double arsR. There are an additional 3 resistance determinants, sul2, strA and strB, combined with Tn3 and IS6 elements.
Comparative analysis between plasmid pLKJULY7989 and pEHS30-1: BLASTn search showed that plasmid pLKJULY7989 is closely associated to pEHS30-1 (accession KX77239.1; 93% coverage; 99% identity) carried by E. coli isolated from human fecal sample, and p1079-IncFIB-N (MG825383.1; 75% coverage; 99% identity) carried by E. coli from chicken stool. The details of comparative analysis were shown in Fig.2.
It is obvious that the length of pEHS30-1 is longer than pLKJULY7989. The pEHS30-1 harbors 15 resistance genes, including aph(3’), sul2, dfrA12, ant(3’), floR, sat1, tetR, tetA, mcr-1 and blaTEM-1. Plasmid pLKJULY7989 and pEH30-1 both possessed the same resistance genes, including sul2, dfrA12, floR, blaTEM-1 and tetR. The middle part of pEHS30-1 (consist of ant (3’), floR, sat1, tetR and tetA) was missing in pLKJULY7989.
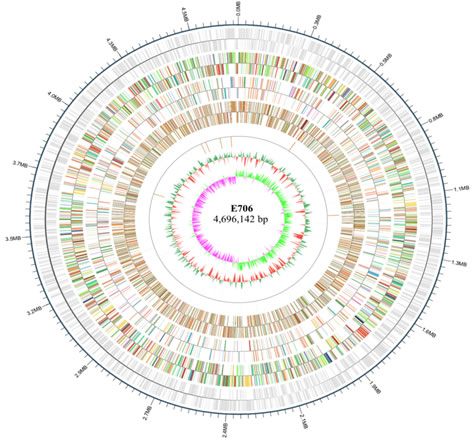
Fig.1 Genome map of Escherichia coli strain (E706)
Rings from the outside as follows: (1) scale marks (unit, Mb), (2) genomic island, (3) protein-coding genes on the forward strand colored by COG category, (4) protein-coding genes on the reverse strand (same color scheme as the second circle), (5) rRNA genes, (6) GC content (deviation from average), and (7) GC skew in blue (below average) and yellow (above average).

Fig.2 Comparative analysis between pLKJULY7989 and pEHS30-1
Open reading frames (ORF) are depicted by arrows. Blue arrows represent genes responsible for replication, yellow arrows are mobile genetic elements, red arrows represent resistance genes, and the degree of genetic similarity between plasmids is depicted by the shaded area.
DISCUSSION
In this study, 271 putative virulence genes were predicted in E706. The strain E706 harbors a variety of virulence genes responsible for the encoding of type IV pilus, biofilm formation and the construction of type IV secretion systems. Besides, E. coli E706 was shown to be mainly resistant to aminoglycoside, macrolide, sulfonamide, anthracyclinones, fosfomycin, quinolones, chloramphenyls, glycopeptide, penicillin, polymyxin, tetracycline, and peptide. Meanwhile, E706 was predicted to be resistant to a variety of heavy metals including As (arsenic), Cu (copper), Mg (magnesium), Mn (manganese), Ni (nickel), Co (cobalt), Ag (silver), Te (tellurium), Zn (zinc), and Cd (cadmium). Recently with the higher use of biocides and heavy metals (Sutterlin et al., 2018), no wondering these resistance genes exist in these isolates. Besides, the coexistence of heavy metal genes or disinfectant genes with antibiotics genes, probably due to the abuse of antibiotics or disinfectant, may lead to co-resistance to these antimicrobials.
The plasmid pLKJULY7989 revealed the fundamental features of the IncFIB backbone encodes genes which is responsible for process of replication, horizontal transfer and stability. To our knowledge, IncF group plasmids have frequently been linked with phenotype of MDR, including PMQR and ESBLs genes (Lyimo et al., 2016; Matsumura et al., 2013).
With the existence of resistance genes, the plasmid pLKJULY7989 could confer resistance to chloramphenicol, penicillins, aminoglycosides and sulfonamide. In addition to antibiotics, the plasmid pLKJULY7989 may also confer resistance to arsenate. The arsR (arsenic regulatory protein), which has high affinity and specificity to arsenite (Ture et al., 2018), is broadly expressed in various species of bacteria, and greatly abundant in few bacterial species (Gao, Yang, & He, 2012). Meanwhile, some regulators of the arsR type have been involved in pathogenesis of bacteria (Gao et al., 2012). Moreover, the amounts of MGEs exist in pLKJULY7989 can also play a vital role in horizontal gene transfer, which would contribute to the dissemination of resistance genes, and offer a promising genetic environment for plasticity of plasmid (Petridis, Bagdasarian, Waldor, & Walker, 2006). An amount of insert sequences such as IS1, IS6 and IS3 have been detected in pLKJULY7989. These sequences are mobile elements which enable the resistance genes to transfer among bacteria. As known, bacteria can acquire resistance genes by the horizontal gene transfer, and integrate genes on the genome to gain acquired resistance depending on these mobile genetic elements (Weng & Zu-Huang, 2013).
It was noteworthy that our plasmid pLKJULY7989 carries class 1 integron with the existence of gene cassette dfrA12-aadA2 and class 1 integrase gene intl1. However, the3’-conserved segment including sul1 (resistance to sulfonamides) and qacEΔ1(quaternary ammonium compound-resistance gene) (Mazel, 2006), was not detected in this plasmid. This finding was consistent with previous study, lacking 3’-conserved region that was generally contained in class 1 integron system (Sáenz et al., 2010).
It is noteworthy that our strain E706 is closely related to strain ECCWS199 and pEHS30-that come from human feces and chicken stool. In our study, the both plasmids pLKJULY7989 and pEHS30-1 are more homologous with the highest coverage of 93% than other plasmids, which probably due to their same IncFIB backbone. They harbor different resistance genes, which confer resistance to different antibiotics. Most compared sequences of both plasmids were predicted as hypothetical proteins. Therefore, the functions of hypothetical protein in pLKJULY7989 should be confirmed in subsequent studies.
In conclusion, we characterized the strain E706, which harbors a series of virulence and resistance genes, CRISPR candidate element, prophage related sequences, methylation modifications as well as genomic islands. Plasmid pLKJULY7989 in E706 is a chimera plasmid made up of a basic IncFIB plasmid backbone with multi-resistance genes and class 1 integron. Plasmid pLKJULY7989 carries class 1 integron with the existence of gene cassette dfrA12-aadA2 and class 1 integrase gene intl1, but lacked the corresponding 3’-conseved segments. The complete sequence will provide valuable information for a better understanding of the genomic structure and pathogenic characteristics of the E. coli.
Nucleotide sequence accession number: The annotated sequence of Escherichia coli strain E706 has been submitted to GenBank under the accession number CP029687 (chromosome) and CP029688 (plasmid).
Acknowledgment: The present study does not cover any studies with human participants executed by any of the authors.
Conflict of interest: No conflict of interest declared.
Funding information: Present study was funded by the “National Natural Science Foundation of China” (31671954), and “Science and Technology Department of Sichuan province” (2020ZHCG0044, 2017JY0118).
REFERENCE
- Blair, J.M., Webber, M.A., Baylay, A.J., Ogbolu, D.O., and Piddock, L.J (2015). Molecular mechanisms of antibiotic resistance. Nat Rev Microbiol. 13(1): 42-51.
- Brettin,T., Davis, J.J., Disz, T., Edwards, R.A., Gerdes, S., Olsen, G.J., and Xia, F (2015). RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep. 5: 8365.
- Carattoli, A., Zankari, E., Garcia-Fernandez, A., Voldby Larsen, M., Lund, O., Villa, L., and Hasman, H (2014). In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob Agents Chemother. 58(7): 3895-3903.
- Colello, R., Kruger, A., Conza, J.D., Rossen, J.A., Friedrich, A.W., Gutkind, G., and Padola, N.L (2018). Antimicrobial Resistance in Class 1 Integron-Positive Shiga Toxin-Producing Escherichia coli Isolated from Cattle, Pigs, Food and Farm Environment. Microorganisms. 6(4): 39-50.
- Flynn, P.B., Higginbotham, S., Alshraiedeh, N.H., Gorman, S.P., Graham, W.G., and Gilmore, B.F (2015). Bactericidal efficacy of atmospheric pressure non-thermal plasma (APNTP) against the ESKAPE pathogens. Int J Antimicrob Agents. 46(1): 101-107.
- Gao, C.H., Yang, M., and He, Z.G (2012). Characterization of a novel ArsR-like regulator encoded by Rv2034 in Mycobacterium tuberculosis. PLoS One. 7(4): e36255.
- Gilrane, V.L., Lobo, S., Huang, W., Zhuge, J., Yin, C., Chen, D., and Wang, G (2017). Complete Genome Sequence of a Colistin-Resistant Escherichia coli Strain Harboring mcr-1 on an IncHI2 Plasmid in the United States. Genome Announc. 5(42).
- Lindsey, R.L., Pouseele, H., Chen, J.C., Strockbine, N.A., and Carleton, H.A (2016). Implementation of Whole Genome Sequencing (WGS) for Identification and Characterization of Shiga Toxin-Producing Escherichia coli (STEC) in the United States. Frontiers In Microbiol. 7: 766.
- Lyimo, B., Buza, J., Subbiah, M., Temba, S., Kipasika, H., Smith, W., and Call, D.R (2016). IncF Plasmids Are Commonly Carried by Antibiotic Resistant Escherichia coli Isolated from Drinking Water Sources in Northern Tanzania. Int J Microbiol. 6(8): 3103672.
- Matsumura, Y., Yamamoto, M., Nagao, M., Ito, Y., Takakura, S., and Ichiyama, S (2013). Association of fluoroquinolone resistance, virulence genes, and IncF plasmids with extended-spectrum-beta-lactamase-producing Escherichia coli sequence type 131 (ST131) and ST405 clonal groups. Antimicrob Agents Chemother. 57(10): 4736-4742.
- Mazel, D (2006). Integrons: agents of bacterial evolution. Nat Rev Microbiol. 4(8): 608-620.
- Palmela, C., Chevarin, C., Xu, Z., Torres, J., Sevrin, G., Hirten, R., and Colombel, J.F (2018). Adherent-invasive Escherichia coli in inflammatory bowel disease. Gut. 67(3): 574-587.
- Pan, Y.S., Zong, Z.Y., Yuan, L., Du, X.D., Huang, H., Zhong, X.H., and Hu, G.Z (2016). Complete Sequence of pEC012, a Multidrug-Resistant IncI1 ST71 Plasmid Carrying blaCTX-M-65, rmtB, fosA3, floR, and oqxAB in an Avian Escherichia coli ST117 Strain. Frontiers In Microbiol. 7: 1117.
- Petridis, M., Bagdasarian, M., Waldor, M.K., and Walker, E (2006). Horizontal transfer of Shiga toxin and antibiotic resistance genes among Escherichia coli strains in house fly (Diptera: Muscidae) gut. J Med Entomol. 43(2): 288-295.
- Runcharoen, C., Raven, K.E., Reuter, S., Kallonen, T., Paksanont, S., Thammachote, J., and Chantratita, N (2017). Whole genome sequencing of ESBL-producing Escherichia coli isolated from patients, farm waste and canals in Thailand. Genome Med. 9(1): 81.
- Sáenz, Y., Vinué, L., Ruiz, E., Somalo, S., Martínez, S., Rojo-Bezares, B., and Torres, C (2010). Class 1 integrons lacking qacEΔ1 and sul1 genes in Escherichia coli isolates of food, animal and human origins. Vet Microbiol. 144(3-4): 493-497.
- Sullivan, M. J., Petty, N.K., and Beatson, S.A (2011). Easyfig: a genome comparison visualizer. Bioinformatics. 27(7): 1009-1010.
- Sutterlin, S., Tellez-Castillo, C.J., Anselem, L., Yin, H., Bray, J.E., and Maiden, M.C (2018). Heavy Metal Susceptibility of Escherichia coli Isolated from Urine Samples from Sweden, Germany, and Spain. Antimicrob Agents Chemother. 62(5).
- Ture, M., Altinok, I., and Alp, H (2018). Effects of Cage Farming on Antimicrobial and Heavy Metal Resistance of Escherichia coli, Enterococcus faecium, and Lactococcus garvieae. Microb Drug Resist. 32(3): 256-263.
- van El, C.G., Cornel, M.C., Borry, P., Hastings, R.J., Fellmann, F., and Hodgson, S.V (2013). Whole-genome sequencing in health care: recommendations of the European Society of Human Genetics. Eur J Hum Genet. 21(6): 580-584.
- Weng, X.B., and Zu, M.I (2013). Mobile genetic elements:carriers of resistance genes. Chinese J Zoonoses. 29(4): 389-397.
- Wong, M.H., Xie, M., Xie, L., Lin, D., Li, R., Zhou, Y., and Chen, S (2016). Complete Sequence of a F33:A-:B- Conjugative Plasmid Carrying the oqxAB, fosA3, and blaCTX-M-55 Elements from a Foodborne Escherichia coli Front Microbiol. 7: 1729.
- Zhang, S., Yang, G., Huang, Y., Zhang, J., Cui, L., and Wu, Q (2018). Prevalence and Characterization of Atypical Enteropathogenic Escherichia coli Isolated from Retail Foods in China. J Food Prot. 7(12): 1761-1767.
- Zhu, M., Yang, G., Li, A., Zong, L., Dong, Z., Lu, J., and Wang, J (2017). Identification and molecular characterization of Escherichia coliblaSHV genes in a Chinese teaching hospital. Gene. 600: 29-35.
|





