ANALYSIS OF CLONED SEQUENCES AND EXPRESSION OF ASR GENE FAMILY IN MILLET
X.N. Cao1,2,3,4,5, L.H. Shen2,3,4, J. Song3,4,5, J.J. Wang3,4,5,H.G. Wang3,4,5· L. Chen3,4,5,Y.X. Pei1 ,S.C. Liu3,4,5* and Z.J. Qiao2,3,4,5*.
1School of Life Science, Shanxi University, Taiyuan, Shanxi 030006, China;
2College of Biological Engineering, Shanxi University, Taiyuan, Shanxi 030006, China; 3Center for Agricultural Genetic Resources Research, Shanxi Agricultural University, Taiyuan 030031 China;4Key Laboratory of Crop Gene Resources and Germplasm Enhancement on Loess Plateau, Ministry of Agriculture, Taiyuan, Shanxi 030031, China; 5Shanxi Key Laboratory of Genetic Resources and Genetic Improvement of Minor Crops, Taiyuan, Shanxi 030031, China
Corresponding author’s email: sxnkyqzj@sina.com; lsch209@163.com
ABSTRACT
ABA-stress-ripening (ASR) protein is a hydrophilic plant-specific protein and plays an indispensable role in plant abiotic stress. However, the study of the ASR gene family of millet has not yet been completed. In this study, we identified six members of the ASR family of millet and analyzed their genetic structure, protein alignment and phylogenetic relationships. The PmASR protein was found to contain highly conserved motifs and the millet ASR gene was divided into three groups, namely (1) PmASR1, PmASR2, PmASR5; (2) PmASR3; (3) PmASR4, PmASR6. Three sets of representative ASR gene clones were selected and the transcriptional profiles of PmASR1, 3, 4 genes in various tissues or under different abiotic stresses were quantitatively analyzed by fluorescence. The results showed that the PmASR1 and PmASR3 genes were mainly up-regulated in roots while PmASR4 gene was mainly up-regulated in leaves. And the expression of ASR genes is different under at least one. Therefore, it can be concluded that the ASR gene family in millet was an abiotic stress response gene.
Key words: Cloning, Millet, ABA-stress-ripening (ASR) protein, Expression patterns
https://doi.org/10.36899/JAPS.2021.5.0332
Published online January 21, 2021
INTRODUCTION
Millet (Panicum miliaceum L.), also known as broomcorn millet and proso millet, is one of the oldest gramineous cereals with a long history of about 8000-10000 years and the major crop in arid and semi-arid regions (Chai, 1999). Millet originated in the Loess Plateau of Yellow River Basin in China and Shanxi is the center of the origin and genetic diversity of millet ( Wang et al., 2009) with the number of germplasm resources ranking first in China (Wang et al., 2018). In addition, as a C4 crop, millet has a higher efficiency of water compared with Millet, Sorghum, Maize and other crops (Wang, 1994). At the same time, the transpiration coefficient of leaves is relatively low while the relative water content and bound water content in leaves are high (Gao et al., 1990; Li, 1996). Moreover, millet, with a strong ability of rehydration, can quickly recover growth after drought. Besides, millet could still accumulate dry matter while C3 crops stopped dry matter accumulation under water stress ( Lin et al., 2006; Huang et al., 1994). Therefore, as an ideal crop to study the mechanism of drought tolerance, millet has attracted more and more attention (Lin et al., 2008; Shi et al., 2019; Zou et al., 2019), which is an excellent material for exploring drought resistance genes and genetic improvement.
ASR (ABA-stress-ripening) genes is mainly responsible for regulating the adaptability of plants to environmental stresses, carbohydrate metabolism, low temperature, trauma, light induction and hormone response in order to maintain the normal activities and development in plants (Wang et al., 2016; Zhang et al., 2018; Liu et al., 2010; Cakir et al., 2003; Liang et al., 2019). At present, ASR gene exists in the form of gene family in plants. There are great differences in ASR gene among different species. It was found that there are 5 ASR genes in tomatoes (Fischer et al., 2011), 7 ASR genes in setavia italica (Feng et al., 2016) and 9 ASR genes in maize (Virlouvet et al., 2011). Besides, the expression patterns and functions of ASR gene family are different even in the same species. For setavia italica, ASR4 and ASR7 are the strongest genes responsing to drought while ASR5 and ASR6 are the strongest genes to salt (Song et al., 2019). The overexpression of ASR1 in tomato can improve its tolerance to drought and salt stress (Amitai-Zeigersona et al., 1995). LLA23 of ASR gene family in lily can improve its tolerance to water and high-salt stress (Yang et al., 2005). At present, there is no research about ASR gene family in millet. In this study, the ASR gene family was cloned from millet and the characteristics were analyzed. In addition, the expressions of ASR genes of different tissues under abiotic stress and hormone stress were detected by qRT-PCR, which laid a foundation for studying the drought resistance and stress tolerance of ASR gene family.
MATERIALS AND METHODS
Materials: The tested material is red millet, which was provided by Shanxi Academy of Agricultural Sciences. The millet seeds were put in 75% ethanol for 2 minutes, soaked in 0.1% mercury for 5 minutes, washed with ddH2O for at least 5 times, and germinated in a germination box with sufficient water in the dark. After germination, the growing buds were put into germination bags under 10,000 LXS light for 16 hours and dark for 8 hours for about 14 days. Then the seedlings were put into 10% PEG6000, 250 mmol/L NaCl, 200 mmol/L mannitol, 100 mmol/L ABA, 100 mmol/L jasmonate methylester, 2 mmol/L salicylic acid, 50 mmol/L auxin, 10 mmol/L hydrogen peroxide and 10 mg/L gibberellin solution for 24 hours. The above solutions of millets represented drought, salt, sugar and plant hormone treatment. Roots, stems and leaves were collected at 3, 6, 9, 12, 24 hours under these treatments and grinded efficiently. Finally, the RNA of millet was extracted by RNAprep pure Plant Kit (DP432 of Tiangen Biotech Co.) according to the instruction manual and frozen in liquid nitrogen.
Identification of ASR family gene in millets: Based on genomic sequences of millet, 36 candidate proteins in total were obtained from NCBI (https://phytozome.jgi.doe.gov/). And the candidating ASR family proteins were selected for CDD alignment in NCBI (http: www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) and used to check whether the conservative domain of ABA-WDS. Finally, six ASR family genes named PmASR1-PmASR6 were obtained.
Bioinformatics analysis: In order to understand the evolutionary relationship of ASR gene, we blasted the ASR protein sequence of millet with multi-sequence alignment of other plants by Clustal X 2.0. The phylogenetic tree was constructed by using the adjacency method (NJ) and bootstrap test in MEGA6 software through 1000 calculations (Hall, 2013). The subcellular localization was analyzed by WoLF PSORT online software. The conservative domain of protein structure was analyzed by Blast in NCBI. The molecular weight, relative molecular weight, isoelectric point and hydrophilicity of the protein were calculated by ExPASy (https://web.expasy.org/compute_pi/). The ASR family gene of millet was analyzed by MEME (http://meme-suite.org/tools/meme).
Cloning of ASR gene in millet: A total of three primer sequences of ASR family gene members were cloned as Table 1. RNA was extracted from the 100 mg collected samples. Then the mRNA was reversely transcribed into cDNA by FastKing one-step method with genomic DNA removed. The template volume was 1μL (100 ng/μL) and the concentration of DNA was 100 ng/μL. A reaction mixture of 20 μL was loaded into an ABI PCR System and subjected to the following cycling: 5 min at 94℃ to denature DNA and activate Taq polymerase; 35 cycles of 5 s at 94℃ and 30s at 58-60℃ and finally at 72℃ to extension and synthesis DNA chain. After detecting the size of the fragments accurately, the target bands were recovered and purified, connected to the cloning vector pGM-Simple-T Fast Vector, transformed into DH5α by heat shock, selected white colonies through blue-white spot screening, and then sent to Shanghai Meijisheng for sequencing. The sequencing results were spliced by DNA MAN software and searched by BLASTN and CD-comparison in NCBI database. Check whether the result contains ABA-WDS conservative domain.
Table 1: Primer sequences
|
Gene
|
Primer
|
Product Length (bp)
|
|
PmASR1
|
Forward:5’-AGAGCGAGCCAAGTGTACTA-3’
Reverse: 5’-CTAAGGGTCCGGAAGGGACT-3’
|
557
|
|
PmASR3
|
Forward: 5’-CTTGTGAGGCAACCCAACAAC-3’
Reverse: 5’-CCGAAGCATGCAGAGTGATTT-3’
|
829
|
|
PmASR4
|
Forward: 5’-CCCCTGACAAGTTCGCAGA-3’
Reverse: 5’-CAGACAGCACCACACGGTTA-3’
|
541
|
qRT-PCR: We detected the mRNA expression of ASR gene under different treatments by qRT-PCR. According to Jiang Xiaomei's report on qRT-PCR internal reference gene of switchgrass (Jiang et al., 2014), ACT2 with high stability under drought, salt and sugar stress was screened as internal reference gene of millet (Table 2). The qRT-PCR reaction system is 20 μL. The reaction procedure was pre-denaturation at 95℃ for 15 min, followed by denaturation at 95℃ for 10 s, annealing and extension at 60℃ for 30 s for 40 cycles.
Table 2: Primers for qRT-PCR
|
Gene
|
Forward
|
Reverse
|
Product Lengths (bp)
|
|
PmASR1
|
5’-ACGAGCACCACGAG
AAGAAG-3’
|
5’-AAGGGTCCGGAAGGGA
CTTA-3’
|
157
|
|
PmASR3
|
5’-GCAGAAGGAGACC
CACTACG-3’
|
5’-TCGTCCCTTCCGATCC
ATCT-3’
|
114
|
|
PmASR4
|
5’-TTGCCTGAACCCTG
AAGCAT-3’
|
5’-AACACCGGACCGAATTC
CTG-3’
|
159
|
|
ACT2
|
5’-GCGAGCTTCCCTG
TAGGTA-3’
|
5’-CGAACCCAGCCTTCAC
CATAC-3’
|
93
|
RESULTS
Identification of ASR family gene and Bioinformatics Analysis of millet: The gene subcellular localization was predicted by WoLF PSORT software, the results showed that the PmASR 1-3 family genes were located in the nucleus. The amino acid sequence, protein molecular weight and subcellular localization of the PmASR gene family were analyzed by ExPASy. The data indicated that the millet ASR proteins were composed of about 100 amino acids. The molecular weight of PmASR gene is between 11-21 KDa and isoelectric point is between 5-10. Except for the predicted results of PmASR5 subcellular in cytoplasm, the others are in nucleus (Table 3).
Table 3: Nomenclature of ASRs in millet
|
Name
|
ORF
|
Amino acids
|
MW(KDa)
|
PI
|
Localization
|
|
PmASR1
|
396
|
131
|
14879.21
|
6.19
|
Nucleus
|
|
PmASR2
|
423
|
140
|
15634.95
|
5.89
|
Nucleus
|
|
PmASR3
|
312
|
103
|
11328.70
|
9.49
|
Nucleus
|
|
PmASR4
|
306
|
101
|
11434.63
|
6.78
|
Nucleus
|
|
PmASR5
|
576
|
191
|
21262.85
|
6.30
|
Cytoplasm
|
|
PmASR6
|
327
|
108
|
11829.10
|
7.34
|
Nucleus
|
We then compared ASR amino acid sequence of millet with other crops (switchgrass, soybean, tomato, setavia italica, maize, wheat, rice, brachypodium distachyon, potatoes and peanuts) containing ASR gene and constructed phylogenetic tree (Fig. 1) with MEGA7.0. These ASR proteins could be divided into 3 groups, accounting for 21%, 21% and 58% of the evolutionary trees for Group1, Group2 and Group3, respectively. And there were 2 ASR proteins (PmASR4 and PMASR6) in Group 1 while Group2 had PmASR3. Group3 was the largest group containing the most ASR proteins, including PmASR1, PmASR2 and PmASR5.Some ASR proteins and dicotyledon ASR proteins were found in the phylogenetic tree. PmASR protein is highly homologous, LyASR protein in tomato and StASR protein in peanut are highly homologous, so ASR gene exists and is very conservative in monocotyledons and dicotyledons.
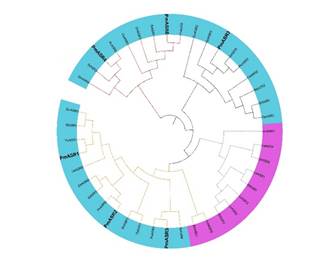
Figure 1. Phylogenetic analysis of ASR proteins in switchgrass, soybean, tomato, setavia italica, maize, wheat, rice, brachypodium distachyon, potato and peanut. And the ASR protein in millet is highlighted by black. The red dotted line was marked as Group 1 while the black dotted line is marked as Group 2, and the others is Group 3. The blue background represents dicotyledons while purple background represents monocotyledons.
Multiple comparisons of ASR family genes (Fig. 2A) and MEME software showed that the similarity of ASR gene ranged from 35.6% to 83.2%. There were highly conserved ABA-WDS conserved
domains at the C-terminal of ASR family genes, which was consistent with the results of protein motif analysis (Fig. 2B). Through phylogenetic tree, we could get PmASR3 and PmASR4; PmASR1 and PmASR2 had high similarities.
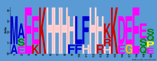
  
Figure 2. Analysis of ASR gene family in millet. A: Phylogenetic tree of the ASR gene family in millet; B: Protein structure analysis of the ASR family in millet; C: Motif of the ASR protein in millet. The background color in C represents as the same meaning as that in B.
Expression of PmASR in different tissues: In order to explore the specificity of the ASR family genes in millet, we analyzed the expression of 6 ASRs in different tissues by qRT-PCR and drew the clustering maps (Fig. 3). It’s easy to find that the expression levels of PmASR1, PmASR2, PmASR3 and PmASR6 were higher in roots while PmASR4 and PmASR5 were mainly expressed in leaves. In addition, PmASR1 was the highest in roots. The expression of PmASR4 was the highest in the leaves among the members. In order to further study the expression analysis of ASR family responding to stress, we took the highest expression of ASR gene PmASR1, PmASR3 and PmASR4 in the roots, stems and leaves of millet, to analyze the expression of ASR gene PmASR1, PmASR3 and PmASR4 under different stress treatments in the phylogenetic tree (Fig. 5&6).

Figure 3. The expression of ASR genes in different parts of millet.The results of qRT-PCR were analyzed by Tanon Gis Tools. Green indicated high level expression while red indicated low level expression. The expression levels of PmASR1, PmASR2, PmASR3 and PmASR6 were higher in roots while PmASR4 and PmASR5 were mainly expressed in leaves. PmASR1 was the highest in roots.
Cloning of ASR gene in millet: We detected the PCR product of millet by 1% agarose gel. The results showed the size of ASR genes as follows: PmASR1:557bp, PmASR3:541bp and PmASR4:829bp (Fig. 4). At the same time, all ASR genes contain ABA/WDS domain with complete coding region by blasting analysis in

NCBI. We predicted the proteins and conserved domains of ASR family in millet by EXPASY software. The results showed PmASR1, 3 and 4 encoded 131, 103 and 101 amino acids, respectively (Table 3), which were rich in glutamic acid, histidine, lysine and other amino acids.

Figure 4. Amplification of ASR gene family in millet. M:DL2000 Marker;1:557bpPmASR1;2:541bpPmASR4;3:829bpPmASR3
Expression of ASR family gene in different tissues under abiotic stress: In order to analyze the expression patterns of PmASR gene family in millet, the expression of PmASR 1, 3 and 4 genes in three tissues under abiotic stress of polyethylene glycol (PEG), NaCl and mannitol were identified by qRT-PCR. The results showed that genes showed different expression patterns under abiotic stress. Among them, the ASRs gene of millet was significantly up-regulated in roots. PmASR 1, 3 and 4 reached a peak at 9h, 3h and 6h (15.7, 8.3, 11.6 times as compared as the control). And they reached the peak at 6h (12.5, 6.7, 3.7 times as compared as the control) in leaf tissues under PEG stress. ASR1 gene was gradually down-regulated in stems while ASR3 was slightly up-regulated, and ASR4 gene did not change. Under NaCl stress, ASR1, 3 and 4 were up-regulated in roots and peaked at 3 h, 9 h and 3 h (9.2, 2.4, 8.0 times as compared as control). At the same time, ASR1, 3 and 4 were up-regulated in leaf and reached the peak values at 3 h, 6 h and 6 h (10.4, 8.7, 3.6 times as compared as control). ASR3 and 4 were up-regulated slightly in stems, and gradually decreased. Under mannitol stress, ASRs were up-regulated, reaching the peak values at 3h, 6h and 3h (5.8, 2.3, 7.2 times as compared as the control), and up-regulated in leaf, reaching the peak values at 3h, 3h and 6h (4.2, 3.3, 5.1 times as compared as control), with little change in stem. These results indicated that the ASR family genes mainly acted on the roots and leaves of millet under osmotic stress.

Figure 5. Expression analysis of ASR gene family in millet under abiotic stress. The mRNA expression differences of PmASR1, 3 and 4 genes in three parts (root, stem and leaf) under abiotic stress of PEG, NaCl and mannitol were analyzed by qRT-PCR. The experiments were repeated at least 3 times. *p<0.05, ** p<0.01.
Expression of ASR family gene in different tissues under phytohormone stress: Because plants often interact with some plant hormones and signaling pathways in response to abiotic stresses, we also examined the effects of abscisic acid (ABA), methyl jasmona (MeJa), salicylic acid (SA), indole-3-acetic acid (IAA), H2O2 and Gibberellin (GA) on PmASRs gene transcription. The ASR1, 3 and 4 genes were up-regulated in roots and leaves under ABA treatment. The ASR1 was up-regulated significantly in roots at 3 h (22.2 times as much as control) while ASR3 was up-regulated most significantly in leaves at 6 h (8.7 times as much as control). Under the treatment of H2O2, the ASR1 gene was up-regulated obviously in the roots at 3 h (12.0 times as much as control), however, there was no significant change in the stems and leaves. ASR3 and 4 were slightly induced in the roots and leaves, but there was no significant change in the stems. Under IAA treatment, the change of ASR1 in roots was the most obvious at 6 h (16.4 times as compared as control) while the changes of ASR3 in stems (6 h) and leaves (9 h) were the most obvious at 6.8 and 11.5 times as compared as control, respectively. Under GA treatment, the ASR1 gene was up-regulated most significantly (8.2 times as much as control) at 3 h in root and 4.7 times in leaf. ASR3 and ASR4 did not change significantly in the roots, stems and leaves under GA treatment. Under SA treatment, ASR1 and ASR4 were up-regulated in root, while ASR3 and ASR4 were up-regulated in leaf. Among them, ASR1 was up-regulated most significantly at 1 h in root (11.9 times as compared as control), and ASR4 was up-regulated most obviously at 3 h in leaf (6.5 times as compared as control). Under MeJa treatment, ASR1 and ASR3 were significantly up-regulated in roots, peaked at 12h and 6h (24.8 and 11.2 times as much as control), slightly up-regulated at 9h (2.2 times as much as control), and up-regulated in leaf, with ASR4 being the most obvious at 9h (6.5 times as much as control).
Figure 6. Expression analysis of ASR gene family in millet under phytohormone stress. The mRNA expression differences of PmASR1, 3 and 4 genes in three parts (root, stem and leaf) under phytohormone stress of ABA, H2O2, IAA, GA, SA and MeJa were analyzed by qRT-PCR. The experiments were repeated at least 3 times. *p<0.05, ** p<0.01.
DISCUSSION
ASR genes, as resistance genes, are mainly involved in osmotic stress regulation among plant species. ASR genes are induced and expressed to protect plant cells from drought, low temperature, salt and other stress ( Amitai-Zeigersona et al., 1995; Henry et al., 2011; Hu et al., 2014). The previous researches showed ASR protein could be used as a molecular chaperone to stabilize cell structure and bind to the excessive metal ions. Besides, ASR protein is also regarded as a transcription factor to regulate the expression of resistance genes (Yang et al., 2005; Kalifa et al., 2004). Therefore, ASR protein is the key bioactive molecules for protecting plants in the course of stress (González et al., 2014). Millet is an important food crop in arid and semi-arid areas of China and there is no report about ASR genes in millet. In this study, we identified 6 ASR genes and cleared the transcription profiles of PmASRs in various tissues by bioinformatics analysis and qRT-PCR. At the same time, we also investigated the expression of PmASR1, PmASR3 and PmASR4 genes under different stress treatments.
Six ASR genes were identified by comparing the genomic sequences in the millet and the known species. By studying the molecular weight, isoelectric point, conservative motif, protein composition and system analysis of the protein, we found the ASR protein in millets was a low molecular weight protein composed of about 100 amino acids, which is similar to that of setavia italica (Feng et al., 2016). The previous study shows that ASR genes in plants usually consists of 2 exons (Henry et al., 2011). Similar conclusions have been drawn in this study. There are 2 exons in PmASR1, PmASR2, PmASR3, PmASR4 and PmASR5. However, there is only 1 exon in PmASR6 in this study., which may be related to the size and complexity of the genome in millet (Huang et al., 2016).
PmASR has a conserved domain of about 100 amino acids at the C-terminal which contains ABA/WDS functional domain (González et al., 2014). Bioinformatics analysis of PmASR showed that the results of subcellular prediction are all in the nucleus except PmASR5 in the cytoplasm (table 3), which indicated that PmASR may play a variety of functions within cells. TaASR1 protein, located in the nucleus in wheat, plays a role of transcription factor and participates the transcription regulation of related genes (Hu et al., 2013). Tomato ASR genes, partly existing in the cytoplasm, has a function of protecting protein. When it binds to DNA in the form of dimer, ASR gene plays a transcription factor transferring from the cytoplasm to the nucleus to reduce the damage on plants (Kalifa et al., 2004).
By analyzing the transcriptional levels of PmASR gene family in different tissues of millet, the results showed that the expression of PmASR4 and PmASR5 were higher in stems and leaves than that in roots, which was consistent with the expression level of ASR genes in mulberry (Zhu et al., 2016). Meanwhile, the expression levels of PmASR1, PmASR2, PmASR3 and PmASR6 were higher in roots, indicating a clear tissue specificity of ASR gene expression.
By studying the expression patterns of PmASR1, PmASR3 and PmASR4 under abiotic stresses (PEG, NaCl and mannitol), the results showed the expressions of PmASR1, PmASR3 and PmASR4 were not significantly changed while the expressions mainly upregulated in roots and leaves. The outcome suggested that ASR gene family mostly acted on the roots and leaves of millet under osmotic stress, and the ASR gene family were closely related to the stress. In addition, ASR gene could regulate Photosynthesis and Respiration, metabolism of carbohydrates, antioxidant system and signaling transduction of plants in order to significantly improve the stress-induced resistance (Amitai-Zeigersona et al., 1995;Yang et al., 2005; Li et al., 2017). Besides, external environment can affect gene expression by DNA methylation. The study on methylation level of ASR1 in tomato leaves reflected the increasing of ASR1 under drought resulted in the number of methylation sites decreasing and the demethylation in intronic regions (González et al., 2011).
The expression of PmASR1 in root and leaves increased significantly under six hormone stresses, especially under MeJa and ABA treatments, which indicated that MeJa has the same effect as ABA treatment on regulating plant metabolism and physiology (Liu et al., 2002). Beyond that, PmASR3 was significantly up-regulated in leaves under ABA and IAA treatments. PmASR4 was especially up-regulated under ABA and SA treatments. SA can reduce the accumulation of radicals and the damage of peroxides on plants under stress (Tao et al., 1999). And the expression of PmASR3 and PmASR4 did not changed obviously under GA treatment. The expressions of PmASR1, PmASR3 and PmASR4 in millet were significantly up-regulated under ABA treatment, which reflected the expression of ASR genes were induced by ABA and ABA signaling pathway existed in PmASR1, PmASR3 and PmASR4. At the same time, the ABA signaling pathway participated in the stress resistance protection of millet. The result was consistent with the ASR gene in wheat , tomato, strawberry and suaeda salsa (Hu et al., 2013;Hu et al., 2014).
To sum up, in this study, six ASR genes are identified in millet. And the expression patterns of ASR genes are different in the roots, stems and leaves. Besides, the ASR gene expressions are clarified under abiotic stresses (PEG, NaCl and mannitol) and hormone stress. Finally, it laid a foundation for further studying the biological function of ASR gene family.
Disclosure Statement: No potential conflicts of interest were disclosed.
Acknowledgement: The work was supported by the Postdoctoral Science Foundation (YCX2018D2BH3), Shanxi Provincial Key Research and Development Program (201603D221003-5,201803D221020-6), the earmarked fund for China Agriculture Research System(CARS-06-13.5-A16), Shanxi Province Science Foundation for Youths (201601D202081), Shanxi Natural Science Foundation (201601D102049) and Shanxi National Academy of Agricultural Sciences National Natural Fund Support and Cultivation Project(Program No. YGJPY2001).
REFERENCES
- Amitai-Zeigersona, H., P. A. Scolnikb and D. Bar-Zvi (1995). Tomato Asr1 mRNA and protein are transiently expressed following salt stress,osmotic stress and treatment with abscisic acid. Plant Science. 110 (2):205-213.
- Cakir, B., A. Agasse, C. Gaillard, A. Saumonneau, S. Delrot. and R. Atanassova1 (2003). A grape ASR protein involved in sugar and abscisic acid signaling. Plant Cell, 15: 2165-2180
- Chai, Y. (1999). Proso millet (Panicum miliaceumL.). Beijing: China Agriculture Press. p1-3 (in Chinese).
- Feng, Z. J., Z.S. Xu, J.T. Sun, L.C. Li, M. Chen, G.X. Yang, G.Y. He and Y.Z. Ma (2016). Investigation of the ASR family in foxtail millet and the role of ASR1 in drought/oxidative stress tolerance. Plant Cell Reports.35(1):115-128.
- Fischer, I., L. Camus-Kulandaivelu, F. Allal and W. Stephan (2011). Adaptation to drought in two wild tomato species: the evolution of the Asrgene family. NewPhytologist. 190: 1032-1044.
- Gao, J.S., R.H. Hu and Y.H. Wei (1990). Osmotic regulation effect of Proso Millet. Nner Mongolia Agricultural Science and Technology. 6: 17-20 (in Chinese).
- González, R.M. and N.D. Iusem (2014). Twenty years of research on Asr (ABA-stress-ripen ing) genes and proteins. Planta.239: 941-949.
- González, R.M., M.M. Ricardi and N.D. Iusem (2011). A typical epigenetic mark in an atypical location: cytosine methylation at asymmetric (CNN) sites within the body of a non-repetitive tomato gene. BMC Plant Biology. 11: 94.
- Hall, B.G. (2013). Building Phylogenetic Trees from Molecular Data with MEGA. Molecular Biology and Evolution.30(5):1229-1235.
- Henry, I.M., S.C. Carpentier, S. Pampurova, A. Van Hoylandt, B. Panis, R. Swennen and S. Remy (2011). Structure and regulation of the Asr gene family in banana. Planta 234:785-798.
- Hu,W., C. Huang, X.M. Deng, S.Y. Zhou, L.H. Chen, Y. Li, C. Wang, Z.B, Ma, Q.Q. Yuan, Y. Wang, R. Cai, X.Y. Liang, G.X. Yang and G.Y. He (2013). TaASR1, a transcription factor gene in wheat, confers drought stress tolerance in transgenic tobacco. Plant Cell & Environment. 36(8):1449-1464.
- Hu,Y.X., X. Yang, X.L. Li, X.D. Yu and Q.L. Li (2014) .The SlASR gene cloned from the extreme halophyte Suaeda liaotungensis K. enhances abiotic stress tolerance in transgenic Arabidopsis thaliana. Gene.549:243-251.
- Huang, K.H., Y. Zhong, Y.J. Li, D. Zheng, and Z.M. Cheng (2016). Genome-wide identifification and expression analysis of the apple ASR gene family in response to Alternaria alternata f. sp. mali.Genome. 59: 866-878.
- Huang,Y.J., Y.q.Jin, D.M. Xu and F.F. Liu (1994).On the Physiological Basis of Anti-adversity and Stable Yield of Millet. Jilin Agricultural Science. 4:18-20(in Chinese).
- Jiang, X. M., X.Q. Zhang, H.D. Yan, Y.Zhang, S.T.Yang and L.K. Huang (2014). Reference gene selection for Real-time quantitative PCR normalization in switchgrass (Panicum virgatum L.) root tissue. Journal of Agricultural Biotechnology.22(1):55-63. (in Chinese).
- Kalifa, Y., A. Gilad, Konrad Z, M. Zaccai, P. Scolnik and D.Bar-Zvi (2004). The water-and salt-stress-regulated Asr1 (abscisic acid stress ripening) gene encodes a zinc-dependent DNA-binding protein. Biochemical Journal. 381:373-378.
- Li, J.R., Y. Dong, C. Li, Y.L. Pan and J.J. Yu (2017). SiASR4, the target gene of SiARDP from Setaria italica, improves abiotic stress adaption in plants. Front Plant Science.7 (12): 2053.
- Li, Q.Q. (1996). Correctly understand the position of broomcorn millet and actively promote high-yield cultivation techniques. Journal of Modern Agriculture. 1: 23-24 (in Chinese).
- Liang, Y.N., Y.L. Jiang, M. Du, B.Y. Li, L. Chen, M.C. Chen, D.M. Jin, J.D.Wu (2019). ZmASR3 from the Maize ASR Gene Family Positively Regulates Drought Tolerance in Transgenic Arabidopsis. International Journal of Molecular Sciences.20, 2278; doi:10. 3390/ijms20092278.
- Lin, F.y., Y.G. Hu, G.Q. Song, H. Zhang, T.M. Liu and B.R. He (2006). Isolation and Analysis of Genes Induced by Rehydration after Serious Drought in Broomcorn Millet (Panicum miliaceum) by Using SSH.Journal of Agricultural Biotechnology.14(4): 537−541(in Chinese).
- Lin,Y.F., S.Q. Wang, Y.G. Hu and B.R. He (2008). Cloning of A S-Adenosylmethionine Synthetase Gene from Broomcorn Millet (Panicum miliaceum L.) and Its Expression during Drought and Re-Watering.ACTA AGRONOMICA SINICA. 34(5):777-782(in Chinese).
- Liu, H.Y., J.R. Dai, D.R. Feng, B. Liu, H.B. Wang and J.F. Wang (2010). Character -ization of a Novel Plantain Asr Gene, MpAsr, that is Regulated in Response to Infection of Fusarium oxysporum f.sp. cubense and Abiotic Stresses. Journal of Integrative Plant Biology. 52 (3):315-323.
- Liu, X., S.Q. Zhang, C.H. Lou (2002). Jasmonic Acid SignalTransduction and It′s Relation to Abscisic Acid Signal Transduction.Plant Physiology Communications. 2002, 38(3): 285-288. (in Chinese).
- Shi, J.P., X.X. Ma, J.H. Zhang, Y.S. Zhou, M.X. Liu, L.L. Huang, S.L. Sun, X.B. Zhang, X. Gao, W. Zhan, P.H. Li, L. Wang, P. Lu, H.M.Zhao, W.B. Song and Jinsheng Lai (2019). Chromosome conformation capture resolved near complete genome assembly of broomcorn millet.Nature Communications. 10:464; https://doi.org/ 10.1038/ s41467 -018-07876-6.
- Song, J., X.N. Cao, H.G. Wang, L. Chen, J.J. Wang, S.C. Liu and Z.J. Qiao (2019).Identification and Expression Analysis of ASR Family Genes in Setaria italica. Crops. (6):33-42. (in Chinese).
- Tao, Z.Y., Q. Zou, T. Peng and S.Y. Cheng (1999). Effects of salicylic acid on osmotic injury on wheat seedling leaves. ACTA BOTANICA BOREALI-OCCIDENTALIA SINICA. 19(2): 296-302. (in Chinese).
- Virlouvet, L., M.P. Jacquemot, D. Gerentes, He´le`ne Corti, S. Bouton, F. Gilard, B. Valot, J. Trouverie, G. Tcherkez, M. Falque, C. Damerval, P. Rogowsky, P. Perez, G. Noctor, M. Zivy, and S. Coursol (2011). The ZmASR1 protein influences branched-chain amino acid biosynthesis and maintains kernel yield in Maize under water-limited conditions. Plant Physiology.157: 917-936.
- Wang, L.Z., W. Hu, J.L. Feng, X.Y. Yang, Q.J. Huang, J.J. Xiao,Y. Liu,G.X. Yang and G.Y. He (2016). Identifification of the ASR gene family from Brachypodium distachyon and functional characterization of BdASR1 in response to drought stress. Plant Cell Reports. 35:1221-1234.
- Wang, X.Y. and L.Wang (2018). Research on the Germplasm Resources of Chinese Proso millet (Panicum miliaceumL.). Beijing: China Agricultural Science and Technology Press. p99-101(in Chinese).
- Wang, X.Y. (1994). Chinese Proso Millet. Beijing:China Agriculture Press.p31-32.(in Chinese).
- Wang, X.Y., L. Wang, Q.F. Wen and Y. Shi. (2009). Shanxi Province is the Center of the Origin and Genetic Diversity of Proso Millet. Journal of Plant Genetic Resources.10 (3):465-470 (in Chinese).
- Yang, C.Y., Y.C. Chen, Y.J. Guang and C.S. Wang (2005). A lily asr protein involves abscisic acid signaling and confers drought and salt resistance in Arabidopsis. Plant Physiology.139(2):836-846.
- Zhang, H., J.X. Zheng, S.G. Jian, K.F. Xia and M.Zhang (2018).Isolation and functional characterization of the ASR gene from Ipomoea pes-caprae.Plant Science Journal,36(3): 402-410. (in Chinese).
- Zhu, P.P., J. Yu, Y.X. Cai, C.Y.Liu, W.B. Wu, C.H. Wang, A.C. Zhao,C. Lu and M.D.Yu (2016) Cloning and Expression Analysis of ASR Gene from Mulberry. Science of Sericulture, 42(1): 0016-0022. (in Chinese).
- Zou C.S., L.T. Li, D. Miki, D.L. Li, Q.M. Tang, L.H. Xiao, S. Rajput, P. Deng, L. Peng, W. Jia, R. Huang , M.L. Zhang, Y.D. Sun, J.M. Hu, X. Fu, P. S. Schnable, Y.X. Chang, F. Li , H. Zhang, B.L. Feng, X.G. Zhu, R.Y. Liu, J. C. Schnable, J.K. Zhu and H. Zhang (2019).The genome of broomcorn millet. Nature Communications.10:436; https:// doi.org/10.1038/s 41467-019- 08409-5.
|





