METAL NANOPARTICLES AFFECT GLOBAL DNA METHYLATION LEVEL AND GENE EXPRESSION IN EARLY GERMINATION OF SOYBEAN
T. M. Linh1*, P. T. Hoe1, N. C. Mai1, N. H. Chau2 and N. T. Van3
1Institute of Marine Biochemistry, 2Institute of Environmental Technology, 3Institute of Biotechnology, Vietnam Academy of Science and Technology, 18 Hoang Quoc Viet, Caugiay, Hanoi, Vietnam.
*Corresponding Author’s E-mail: tmlinh@imbc.vast.vn
ABSTRACT
Nowadays, metal-based nanoparticles have been widely applied at a small dose for plant-growth and agronomic yield promotion, leading to significant economic and environmental benefits. The interaction mechanism between nanoparticles (NPs) and plants has been reported, however, not been completely elucidated yet. Therefore, this study aims to look at the global DNA methylation status and the expression of germination- related genes of NP treated soybean plants. The analysis indicated that the global DNA methylation, one important epigenetic mechanism in controlling gene expression, of NP treated plants was obviously changed from the control plants, especially in primary roots. The DNA methylation levels in primary roots were decreased significantly at 18 hours and increased at 48 hours after seed treatment with Fe or Co NPs and these values were opposite with Cu, ZnO NP treatment. Together with changes in DNA methylation, the expression of representative germination-related genes such as Glyma.08G018000, Glyma.07G007000, Glyma.09G21700, Glyma.04G096500 and Glyma.02G163900, which encoding for enzymes involved in ethylene biosynthesis and mobilization of seed reserves was increased under all NP treatments. The obtained results suggest that metal-based nanoparticles stimulate seed germination and seedling growth of soybean via regulation of DNA methylation and related gene expression.
Key words: DNA methylation, gene expression, germination, metal nanoparticles, seed treatment, soybean.
https://doi.org/10.36899/JAPS.2021.5.0347
Published online January 21, 2021
INTRODUCTION
Mineral nutrition is an indispensable requirement for the increase in crop productivity, especially in case of growing population worldwide. Recently, nanotechnology is known to provide solutions to enhancing agricultural productivity with limited inputs due to transformation to new biologically forms (Morales-Díaz et al., 2017). Mineral nanoparticles (NPs), both major essential (K, Ca, Mg, Na, P and S) and essential trace (Fe, Mn, Zn, Cu, B, Mo, As, Se, Ni, V, Cr and Co) have been reported to affect plant physiology and development, depending on their sizes, composition, concentration and mode of application. ZnO NPs at various concentrations have been showed to enhance the germination and plant growth in peanut Arachis hypogaea (Prasad et al., 2012), Cicer arietinum (Pandey et al., 2010) and Zea mays (Adhikari et al., 2015). Fe2O3 NPs also have a similar role as ZnO NPs in improving germination, young plant growth, and antioxidant enzymes in Citrullus lanatus (Li et al., 2013). The growth and glycoprotein and protein concentrations in leaves of A. hypogea also found to be promoted when seeds were treated with another Fe NP form - Fe3O4 at a concentration of 4000 mg/L (Suresh et al., 2016). The vigor, synthesis and mobilization of starches and proteins in seedlings of Z. mays were increased in case seeds priming with Cu-chitosan NPs (Saharan et al., 2016). Our recent study also found the positive effects of Fe, Cu, ZnO and Co NPs on germination of soybean (Pham et al., 2018).
The physiological and molecular mechanisms behind nano-mineral effects on plants have been reported, however, there are many questions remained to be addressed. Lahiani et al. (2013) was only one of few studies dealing with molecular mechanism of soybean, maize and barley seeds responded with NPs and revealed that multi-walled carbon nanotubes (MWCNTs) had a role in regulating the expression of genes encoding several types of water channel proteins. Khodakovskaya et al. (2012) suggested that MWCNTs could act as regulators for seed germination and plant growth by boosting the expression of CycB, NtLRX1 and NtPIP1 which were considered as the marker genes for cell division, cell wall formation and water transport, respectively.
Different from animals, plants cannot run away from environmental stresses. Therefore, the developmental plasticity is required and the flexible epigenetic regulation is essential for reprogramming of plant gene expression for rapid responses and adaptation to environmental cues. The information regarding DNA methylation state in plants provides important knowledge of the gene expression control. During different stages of seed maturation of soybean, An et al. (2017) found that DNA methylation levels in promoter regions were dynamics. In different organs of tomato or rice and Silene latifolia, significant alterations of DNA methylation level have been observed (Messeguer et al., 1991; Xiong et al., 1999; Zluvova et al., 2001). The variation in DNA methylation even existed within a population with a similar genotype such as among 96 natural accessions of Arabidopsis thaliana (Vaughn et al., 2007). Our previous results (Pham et al., 2018), in which the improvement in soybean germination under NP treatments was obtained, led us to speculate that the better germination performance of soybean may be consequences of DNA de-methylation and expression enhancement of enzymes involved in the soybean germination process. Han et al. (2013) reported the expression of several proteins involving lipid, protein and carbohydrate metabolism in soybean germinating seeds. Most of lipid metabolism- and amino acid metabolism- related proteins are belong to the lipoxygenase family and the usease, methionine synthase, respectively. In addition, ten groups of carbohydrate metabolism-related proteins were detected, including- amylase and α-glucan water dikinase (Han et al., 2013). Recently, a transcriptome analysis of soybean embryonic axes revealed that a number of genes involved in the biosynthesis and signaling of the germination-promoting hormones including gibberellins and ethylene were up-regulated in different stages during germination (Bellieny-Rabelo et al., 2016).
In practice, the improvement of plant development traits including seed germination under NP application was not inherited into the next generation, but repeated use of NP has to be done every season. Therefore, the NP effects may be associated with epigenetic regulation such as global DNA methylation level and epigenetic expression of key genes, which were known to modulate physiology and development of plants. In order to elucidate the possible roles of NPs in stimulating soybean germination and young plant growth, the global DNA methylation level of NP treated soybean seeds during germination was investigated in this study. Following, the semi quantitative expression of representative genes encoding for 1-aminocyclopropane-1-carboxylic acid synthase (ACC synthase) in ethylene biosynthesis, lipoxygenase in lipid mobilization, urease in nitrogen metabolism, β- amylase and α-glucan water dikinase in carbohydrate metabolism was also examined. The obtained results could partly set a light in the effect mechanism of NPs in germination and development of plants and provide knowledge for confident and sustainable application of nano-materials in agriculture production.
MATERIALS AND METHODS
Seed treatment: Seeds of soybean cultivar DT26 harvested in the winter of 2017 were provided by Legumes Research and Development Center, Food Crops Research Institute, Vietnam Academy of Agriculture Sciences. Healthy and unique size seeds were selected and surface sterilized using chlorine gas as described by Clough and Bent (1998). Iron (Fe), zinc oxide (ZnO), copper (Cu) and cobalt (Co) nanoparticles with purity > 95% and 40-60 nm in size, provided by the Institute of Environmental Technology, Vietnam Academy of Science and Technology (VAST), were suspended in distilled deionized water by ultrasonic at 200 W and 37 kHz for 30 min. Sterilized seeds (150 seeds, app. 30 g) were soaked for 30 min in 10 mL of 50 mg/L Fe NP; 50 mg/L ZnO NP; 50 mg/L Cu NP or 0.05 mg/L Co NP, separately. These concentrations are chosen according to obtained results from our previous study (Pham et al., 2018). Control was treated with 10 mL of distilled deionized water. Seeds (50 per replication) were germinated on 10 x 20 x 15 cm rectangular plastic containers containing 0.5% agar solution (Duchefa Biochemie BV, catalog number: D1001) in darkness at room temperature. Each NP treatment was performed in three replications.
For DNA and RNA analysis, primary root and leave samples were collected at 18, 48 hours (h) and 63, 87 h after NP treatment, respectively. The collected samples were immediately put in liquid nitrogen and kept at -80°C for further use. All experiments were carried out at Institute of Marine Biochemistry, VAST in 2018.
DNA extraction and digestion: Total genomic DNA was extracted from plant samples using cetyl trimethylammonium bromide (CTAB) method (Doyle, 1991). RNA contamination was removed by addition of 1 µL RNase A 10 mg/mL and incubation at 37oC for 2 h. After digestion, 1 µL of EDTA 50 mM was added and incubated at 65oC in 10 min, followed by 3 M NaOAc (pH 5.4) to a final concentration of 0.3 M. DNA was precipitated with ice-cold isopropanol 100% for 30 min at -20oC and were collected by centrifuging at 13000 rpm for 15 min, 4oC. The DNA pellet were washed with 0.5 mL of 70% (v/v) cold ethanol, air dried, dissolved in 30 µL of deionized free DNase RNase H2O and stored at -20oC. DNA purity and concentration were estimated using Nanodrop spectrophotometer (Thermo Scientific).
Double stranded DNA was denatured by incubating at 95oC for 5 min and cooled immediately on ice for 5 min. DNA samples (40 µg DNA in 20 µL of deionized H2O) were digested by adding 2 µL of nuclease P1 solution (0.1 U/µL nuclease P1 in 30 mM NaOAc pH 5.2; 5 mM ZnCl2 and 50 mM NaCl) and incubating 37oC for 4 h. Following the digestion, 0.5 µL of alkaline phosphatase (0.1 U/µL) and 2.5 µL of 10X dephosphorylation buffer was added. Then, the solution was mixed gently and incubated at 37°C for 4 h. The digested DNA was collected by centrifuged at 10000 rpm for 5 min and stored at -20oC until HPLC analysis.
DNA methylation analysis: Plant DNA methylation analysis was performed using HPLC method as recommended by Johnston et al. (2005) and Magana et al. (2007) with minor modifications suitable for our samples and HPLC system. The modifications are as following: 10 µL of DNA sample (0.45 µm filtered) (equivalent to 15 µg of hydrolysed DNA) was injected onto a HPLC Agilent 6120 Series Quadrupole LC/MS System with VertiSep GES C18 column (4.6 x 150 mm, 5 μm particle size) attached to a VertiSep GES C18 guard column (4.6 x 10 mm, 5 μm particle size) (Vertical, Thailand). The gradient program for this column: 0-5 min – 5%A, 10-15 min- 10%A, 25 min-50%A and 30-50 min- 95%A at a total flow rate of 0.5 mL/min for 50 min at 25°C (solvent A, MeOH and solvent B, H2O). The results monitored at 275 nm with UV detection. The calibration lines for 5-methyl-deoxy-cytidine (5mdC) and deoxy-cytidine (dC) were constructed by the dilution of stock standard solution (2 mg/mL) for 5mdC (0.5-50 µg/µL) and dC (1-200 µg/µL). The level of 5mdC present in the DNA samples was calculated using the following equation:
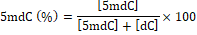
In which [5mdC] and [dC] are the respective concentration of 5-methyldeoxycytidine and deoxycytidine, as deduced from the calibration curves for external standards of known concentrations.
Each analysis was performed with three biological replicates and experimental data was statistically analysed using Microsoft Excel software. The data were analysed using one-way ANOVA followed by Tukey Test. Differences between the means were considered significant at p<0.05.
Reverse transcription polymerase chain reaction analysis: Gene information and sequences were obtained from National Center for Biotechnology Information -NCBI (https://www.ncbi.nlm.nih.gov/) and Phytozome v11 database (https://phytozome.jgi.doe.gov/pz/portal.html) (Goodstein et al., 2012). Specific PCR primer pairs for each gene were designed using Primer3Plus software (http://frodo.wi.mit.edu/primer3/). The sequences of primers used are listed in Table 1.
Table 1: Gene information and primer sequences for PCR analysis.
|
No.
|
Gene ID
|
Gene description
|
Primer sequences (5’-3’)
|
Amplicon length (bp)
|
|
1.
|
Glyma.08G018000
|
1-aminocyclopropane-1-carboxylic acid synthase
|
GTAGAACAAGAGGAAACAGAGTC
|
389
|
|
AGATACAAGATGGATACGCTTC
|
|
2.
|
Glyma.03G237300
|
Linoleate 9S-lipoxygenase 5
|
GCGTGCTTCACCCTATTTAT
|
360
|
|
CTGTGTCATCTTCCTTGTAGTAG
|
|
3.
|
Glyma.07G007000
|
Lipoxygenase
|
GAAGATGTGCGTAGTCTCTATGA
|
424
|
|
CAGTAGGTAGGCTGTTTATCCT
|
|
4.
|
Glyma.09G21700
|
β-amylase 8
|
GTATCCTGGTATCGGTGAGTT
|
450
|
|
CTGGAAGATGAAATCCTAAGC
|
|
5.
|
Glyma.05G219200
|
β- amylase 3
|
GTTTACGATTGGAGAGGGTATG
|
388
|
|
CCTGTGAAGAGAATGAAGGATA
|
|
6.
|
Glyma.04G096500
|
α-glucan water dikinase 2
|
GGAATAGCTATCTAACTGGAGGA
|
424
|
|
CTATTCCCTGCAAAGTTATCTC
|
|
7.
|
Glyma.02G163900
|
Urease accessory protein UreD
|
GACGGTAGAGGAAGAATAATGAG
|
292
|
|
TGTAAACCTTAGTGGAACCTTG
|
|
8.
|
Glyma.14G039400
|
Urease accessory protein UreF
|
GTTCAGTCCCACTTAGTGTCTAAT
|
292
|
|
ATACAGTCTCTAAACCCAGAGAAG
|
|
9.
|
Glyma.19G147900
|
Actin
|
GACCTTCAACACCCCTGCTA
|
423
|
|
ATGGATGGCTGGAACAGAAC
|
Total RNA was isolated using TRizol reagents (Rio et al., 2010) and then treated with RNase-free DNaseI 1 U/µL (Thermo Scientific) to remove residual genomic DNA. First-strand cDNA was synthesised from 1 µg RNA using Revert Aid First Strand cDNA Synthesis Kit (Thermo Scientific) as the manufacturer’s guidelines. PCR was carried out in a 25 µL reaction containing 1 µL cDNA, 2.5 µL 10X PCR buffer, 2 µL dNTP (2.5 mM), 2 µL MgCl2 (25 mM), 0.5 µL Primer F (10 µM) and 0.5 µL Primer R (10 µM ), 0.5 µL Dream Taq polymerase (5 U/µL) and 16 µL H2O. Thermal cycling began with a denaturation step at 95oC for 10 min, followed by 35 cycles at 95oC for 45 seconds, 54oC for 45 seconds and 72oC for 1 min, and a final extension at 72oC for 7 min. The PCR products were identified on 0.8% agarose gel electrophoresis stained with ethidium bromide.
RESULTS
DNA methylation analysis of germinated soybean from NP treated seeds: In this study, DNA methylation of germination seeds treated with Fe, Cu, ZnO and Co NP was analysed in order to elucidate the possible molecular mechanism associated with morphological responses during early germination of NP treated soybean seeds, which was obtained in our previous report (Pham et al., 2018). The primary roots and leaves of germinated soybean seeds were collected from 18 and 63 h after NP treatment, respectively, because it was the time they can be obviously seen.
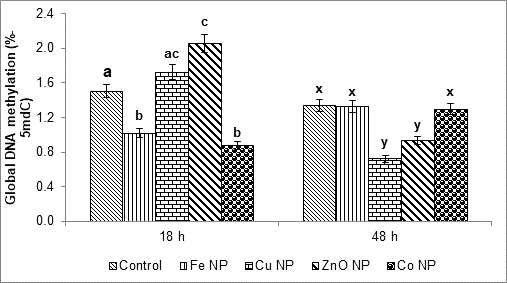
Figure 1: The effect of metal NPs on DNA methylation levels of primary roots at 18 and 48 h after NP treatments. Values are the means of three replicate samples and bars indicate standard errors. Means with different letters are significant different (Tukey’HSD, p ≤ 0.05).
The degree of DNA methylation is expressed as a percentage of the 5mdC in the analysed DNA samples. The DNA methylation levels in primary roots of the NP treated seeds were alternated significant compared to the control (Figure 1). At 18 h (when the primary roots ~ 10 mm in length), the DNA methylation level in primary roots of Fe and Co NP treated seeds decreased significantly compared to the control, from 1.53% down to 1.02 and 0.87%, respectively. Meanwhile, the DNA methylation levels with Cu and ZnO NP treatments were increased up to 1.72% and 2.06%, respectively. After 48 h from NP treatment (when root lengths reaching ~ 30 mm), the DNA methylation levels in primary roots of the Fe and Co NP treated seeds became similar to the control. In opposite to those at 18 h time point, DNA methylation levels in primary roots of the Cu and ZnO NPs at 48 h time point were significantly lower than the control, from 1.34% down to 0.93 and 0.72%, respectively (Figure 1).
Different from primary roots, the DNA methylation levels in primary leaves of NP treated seeds at 63 h were higher than the control, especially with Cu and ZnO NP treatments. Their DNA methylation levels were up significantly from 1.2% 5mdC of the control to 1.88 and 1.56% 5-mdC, respectively. At 87 h, DNA methylation levels of all NP treatments return to the control level (Figure 2).
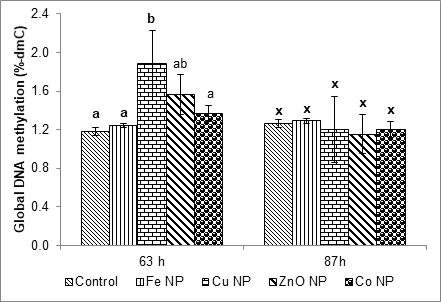
Figure 2: The effect of metal NPs on DNA methylation levels of primary leaves at 63 and 87 h after NP treatments. Values are the means of three replicate samples and bars indicate standard errors. Means with different letters are significant different (Tukey’HSD, p ≤ 0.05).
PCR analysis of germination related genes: The treatment of soybean seeds with metal NP before sowing has a significant influence on the expression of 8 representative germination-related genes in primary root at both time points of 18 and 48 h after treatment. Although, expression of the housekeeping gene (Actin, Glyma.19G147900) was equal among samples, the expression levels of targeted genes seem to depend on NP types and development periods (Figure 3).
The results shown in Figure 3 reveal that, most of studied genes (except Glyma.14G039400) did not transcript in 18-hour- primary roots without NP treatment since no PCR product was observed in the control primary roots (Ctrl lane). Meanwhile, five genes, including Glyma.08G018000 (encoding for ACC synthase), Glyma.07G007000 (encoding for lipoxygenase), Glyma.09G21700 (encoding for beta-amylase), Glyma.04G096500 (encoding for glucan water dikinase) and Glyma.02G163900 (encoding for urease), were found to express under all NP treatments. Among them, the strongest were Glyma.08G018000, Glyma.07G007000 and Glyma.09G21700, following Glyma.02G163900 and Glyma.04G096500. The two genes, Glyma.03G237300 and Glyma.05G219200 were only expressed under Fe and ZnO NP treatments.
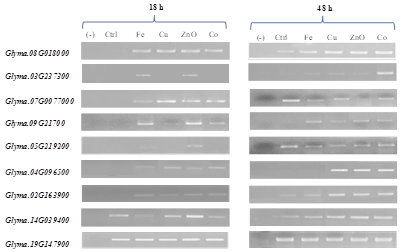
Figure 3: The expression of representative germination-related genes in primary roots at 18 and 48 h after NP treatments. (-): negative control of PCR (without template); Ctr: primary roots from untreated seeds; Fe, Cu, ZnO and Co: primary roots from NP treated seeds.
The above pattern was changed at the time point of 48 h. Apart from the house keeping gene (Glyma.19G127900), the expression of Glyma.08G018000, Glyma.07G007000, Glyma.05G219200, Glyma.02G163900 and Glyma.14G039400 was observed in primary roots for all treatments, including the control. While Glyma.03G237300 and Glyma.09G21700 genes were found to be expressed under all 4 NP treatments, Glyma.04G096500 gene was only expressed in primary roots under Cu, ZnO and Co NP treatments. The expression of Glyma.03G237300 was seen due to NP treatments; however, it was the strongest with Co NP treatment (Figure 3).
In primary leaves, NP treatments make very little changes in the expression of interested genes at the early stages of soybean germination (63 h and then 87 h after NP treatments) (Figure 4). This obtained result is also matching with the DNA methylation levels of primary leaves shown in Figure 2.
Figure 4: The expression of representative germination-related genes in primary leaves at 63 and 87 h after NP treatments. (-): negative control of PCR (without template); Ctr: primary leaves from untreated seeds; Fe, Cu, ZnO and Co: primary leaves from NP treated seeds.
DISCUSSION
Gene expression was known to be regulated by both epigenetic and genetic mechanisms, in which epigenetic regulation mechanisms at molecular level include DNA methylation, histone modifications, and RNA-based mechanisms (Chen et al., 2010). In this study, the DNA methylation during the soybean seed germination under NP treatment was analysed using HPLC method, which is considered to be the gold standard for quantifying methylated cytosine. The changes in global DNA methylation levels within hours of germination could be coupled with the enhancement in the growth rate of primary roots and leaves under NP treatments. While NPs seem have a little effect on primary leaf development at early stages, DNA de-methylation was observed in primary roots after 18 and 48 h from NP treatments. This phenomenon may generate more or higher expression of genes needed for germination process. The DNA methylation is known to be dynamic during plant development and its levels in gene promoters can change during different stages of soybean development (An et al., 2017). When compare the DNA methylation levels of dry and germinating Arabidopsis seeds, Kawakatsu et al. (2017) observed the reprogramming of global DNA methylation during both phases and gave an explanation that the high level DNA methylation in dry seeds as results of preventing the expression of unfavorable genes or transposable element activation. In opposite, this level could be reduced to promote the expression of germination-related genes when conditions for germination are available.
Several genes encoding to germination-related enzymes were selected for examination their expression in germinated soybean seeds after NP treatments. Among them, eight genes, Glyma.08G018000, Glyma.03G237300, Glyma.07G007000, Glyma.09G21700, Glyma.05G219200, Glyma.04G096500, Glyma.02G163900, Glyma.14G039400 have been changed their expression between NP treatments and the control. The results here revealed that most above genes showed different expression pattern at earlier stage (18 h) between NP and non-NP treatment (control) primary roots. At this time point, the expression of most genes tends to be enhanced in NP-treated seeds. The expression of Glyma.08G018000 encoding for ACC synthase was presented earlier in primary roots (at 18 h time point) and seemed to be increased under all NP treatments (at 48 h time point). According to Corbineau et al. (2014), ethylene is involved in the boost of germination of non-dormant seeds. Therefore, its production in non-dormant seeds was found much higher than in dormant ones (Matilla, 2000; Corbineau et al., 2014) and positively correlated with the germination rate of non-dormant seeds (Gorecki et al., 1991). The highest amount of ethylene was noticed when germination process was completed, however it was already measureable long before radicle through seed coat. The expression of ethylene biosynthesis enzymes such as 1-aminocyclopropane-1-carboxylic acid oxidase (ACO, the ethylene-forming enzyme) during seed germination was reported in legume species (Petruzzelli et al., 2000).
Similar to Glyma.08G018000, mRNA expression of Glyma.03G237300 and Glyma.07G007000 encoding for lipoxygenases (LOXs) was increased in primary roots, especially at 18 h time point. LOXs are normally present in the seeds of plants and should be abundant in soybean seed, where lipids are known to be one of major reserves. During germination, the levels of LOX enzyme were observed to be increased (Uma et al., 2005). Maximal accumulation of LOX protein and the corresponding mRNAs lasted from a few hours to few days after germination. With a proteomic analysis of germinating soybean seeds, Han et al. (2013) found the present of a number of LOX enzymes, suggesting the oils in soybean seeds might be degraded through a LOX-dependent pathway. The presence of LOX in the germination process was also detected in pea seeds (Pisum sativum L) (Mo and Koster, 2006).
The increased expression of genes for amylase may accelerate seed germination. According to Mahakham et al. (2017) the improvement of the germination ability of rice seeds treated with metal NPs was consequences of the increase in α-amylase activity and the water absorption by enhancing expression of genes coding for water channel proteins (aquaporin). Our result also showed the early expression of Glyma.09G21700 and Glyma.05G219200 to produce β-amylases at the 18 h in germinating seeds treated with NPs. This observation correlated with the fact that amylases play a key role in the mobilization of energy reserves stored in insoluble starch.
However, different from other crops, stored reservation of soybean mainly contains oils and proteins, the mobilization of reserves during germination including lipids and proteins, not only starches. Urease has been proposed by Thompson (1980) to play an important function in the utilization of seed protein reserves during germination. It can convert urea to ammonia for nitrogen assimilation (Wang et al., 2008) and promoted the germination of nitrogen-limited Arabidopsis thaliana seeds (Zonia et al., 1995). Recently, urease also was reported as one of the main group of amino acid metabolism-related proteins found in germinating soybean seeds (Han et al., 2013). In this study, the increase in expression of Glyma.02G163900 encoding for urease could lead to activating protein mobilization in germinating soybean germination. Altogether, the high expression level of studied enzymes in germinating NP treated-seeds could partly explain for the positive effect of NP treatments in germination improvement of local DT26 soybean cultivar as described in our previous study (Pham et al., 2018).
Conclusion: There are evidences from our studies that the NP treatments of seeds before sowing trigger DNA methylation pattern changed in germinated soybean, especially in primary roots. That may lead to the expression enhancement of important germination-related genes and as consequence to promote germination process of experimental soybean cultivar. However, quantitative information about related gene expression and protein accumulation should be done in different development stages of soybean plants in order to confident application of NP seed treatment for soybean production.
Acknowledgements: This work was financially supported by Vietnam Academy of Science and Technology for the key project “Application of Nanotechnology in Agriculture” project code: VAST.TĐ. NANO-NN/15-18. Thanks go to the Core Facility of Institute of Marine Biochemistry for HPLC analysis. We also thank Dr. Ngo Quoc Buu and Mr. Dao Trong Hien (Institute of Environmental Technology) for NP preparation.
REFERENCES
- Adhikari, T., S. Kundu, A. Biswas J. C. Tarafdar, and A. Subba Rao (2015). Characterization of zinc oxide nano particles and their effect on growth of maize (Zea mays) plant. J. Plant Nutr. 38: 1505-1515.
- An, Y.C., W. Goettel, Q. Han, A. Bartels, Z. Liu, and W. Xiao (2017). Dynamic Changes of Genome-Wide DNA Methylation during Soybean Seed Development. Sci. Rep. 7(1): 12263.
- Bellieny-Rabelo, D., E. Alves Gamosa de Oliveira, E. da Silva Ribeiro, E. Pessoa Costa, A. Elenir Amâncio Oliveira, and T. Motta Venancio (2016). Transcriptome analysis uncovers key regulatory and metabolic aspects of soybean embryonic axes during germination. Sci. Rep. 6: 36009.
- Chen, M., Lv. Shaolei, and Y. Meng (2010). Epigenetic performers in plants: Review of plant epigenetics. Dev. Growth Differ. 52(6): 555-566.
- Clough, S.J. and A.F. Bent (1998). Floral dip: A simplified method for Agrobacterium mediated transformation of Arabidopsis thaliana. Plant J. 16: 735-743.
- Corbineau, F., Q. Xia, C. Bailly, and H. El-Maarouf-Bouteau (2014). Ethylene, a key factor in the regulation of seed dormancy. Front. Plant Sci. 5: 539.
- Doyle, J. (1991). DNA Protocols for Plants. In: Hewitt G.M., Johnston A.W.B., Young J.P.W. (eds) Molecular Techniques in Taxonomy. NATO ASI Series (Series H: Cell Biology), vol 57. Springer, Berlin, Heidelberg. pp: 283-293.
- Goodstein, D.M., S. Shu, R. Howson, R. Neupane, R.D. Hayes, J. Fazo, T. Mitros, W. Dirks, U. Hellsten, N. Putnam, and D.S. Rokhsar (2012). Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40: 1178-1186.
- Gorecki, R.J., H. Ashino, S. Satoh, and Y. Esashi (1991). Ethylene production in pea and cocklebur seeds of differing vigour. J. Exp. Bot. 42(3): 407-414.
- Han, C., X. Yin, D. He, and P. Yang (2013). Analysis of proteome profile in germinating soybean seed, and its comparison with rice showing the styles of reserves mobilization in different crops. PLoS One. 8(2): e56947.
- Johnston, J.W., K. Harding, D.H. Bremner, G. Souch, J. Green, P.T. Lynch, B. Grout, and E.E. Benson (2005). HPLC analysis of plant DNA methylation: a study of critical methodological factors. Plant Physiol. Biochem. 43: 844-853.
- Kawakatsu, T., J.R. Nery, R. Castanon, and J.R. Ecker (2017). Dynamic DNA methylation reconfiguration during seed development and germination. Genome Biol. 18(1): 171.
- Khodakovskaya, M. V., K. De Silva, A.S. Biris, E. Dervishi, and H. Villagarcia (2012). Carbon nanotubes induce growth enhancement of tobacco cells. ACS Nano. 6(3): 2128–2135.
- Lahiani, M.H., E. Dervishi, J. Chen, Z. Nima, A. Gaume, A.S. Biris, and M.V. Khodakovskaya (2013). Impact of carbon nanotube exposure to seeds of valuable crops. ACS Appl. Mater. Interfaces. 5(16): 7965-7973.
- Li, J., P.R. Chang, J. Huang, Y. Wang, H. Yuan, and H. Ren (2013). Physiological effects of magnetic iron oxide nanoparticles towards watermelon. J. Nanosci. Nanotechnol. 13(8): 5561-5567.
- Magana, A.A., K. Wrobel, Y.A. Caudillo, S. Zaina, G. Lund, and K. Wrobel (2007). High-performance liquid chromatography determination of 5-methyl-2'-deoxycytidine, 2'-deoxycytidine, and other deoxynucleosides and nucleosides in DNA digests. Anal. Biochem. 374(2): 378-385.
- Mahakham, W., A.K. Sarmah, S. Maensiri, and P. Theerakulpisut (2017). Nanopriming technology for enhancing germination and starch metabolism of aged rice seeds using phytosynthesized silver nanoparticles. Sci. Rep. 7(1): 8263.
- Matilla, A.J. (2000). Ethylene in seed formation and germination. Seed Sci. Res. 10(2): 111-126.
- Messeguer, R., M.W. Ganal, J.C. Steffens, and S.D. Tanksley (1991). Characterization of the level, target sites and inheritance of cytosine methylation in tomato nuclear DNA. Plant Mol. Biol. 16(5): 753-770.
- Mo, B. and K.L. Koster (2006). Changes in lipoxygenase isoforms during germination and early seedling growth of Pisum sativum. Seed Sci. Res. 16: 97-106.
- Morales-Díaz, A.B., H. Ortega-Ortíz, A. Juárez-Maldonado, G. Cadenas-Pliego, S. González-Morales, and A. Benavides-Mendoza (2017). Application of nanoelements in plant nutrition and its impact in ecosystems. Adv. Nat. Sci: Nanosci. Nanotechnol. 8(1): 013001.
- Pandey, A.C., S.S. Sanjay, and R.S. Yadav (2010). Application of ZnO nanoparticles in influencing the growth rate of Cicer arietinum. Exp. Nanosci. 5(6): 488-497.
- Petruzzelli, L., I. Coraggio, and G. Leubner-Metzger (2000). Ethylene promotes ethylene biosynthesis during pea seed germination by positive feedback regulation of 1-amonocyclopropane-1-carboxylic acid oxidase. Planta. 211(1): 144-149.
- Pham, T.H., N.C. Mai, L.Q. Lien, N.K. Ban, C.V. Minh, N.H. Chau, N.Q. Buu, D.T. Hien, N.T. Van, L.T.T. Hien, and T.M. Linh (2018). Germination responses of soybean seeds under Fe, ZnO, Cu and Co nanoparticle treatments. Int. J. Agric. Biol. 20(7): 1562-1568.
- Prasad, T.N.V.K.V., P. Sudhakar, Y. Sreenivasulu, P. Latha, V. Munaswamy, K. Raja Reddy, T.S. Sreeprasad, P.R. Sajanlal, and T. Pradeep (2012). Effect of nanoscale zinc oxide particles on the germination, growth and yield of peanut. J. Plant Nutr. 35(6): 905-927.
- Rio, D.C., M. Ares, G.J. Hannon, and T.W. Nilsen (2010). Purification of RNA using TRIzol (TRI reagent). Cold Spring Harbor Protocols.
- Saharan, V., R.V. Kumaraswamy, R.C. Choudhary, S. Kumari, A. Pal, R. Raliya, and P. Biswas (2016). Cu-Chitosan nanoparticle mediated sustainable approach to enhance seedling growth in maize by mobilizing reserved food. J. Agric. Food Chem. 64(31): 6148-6155.
- Suresh, S., S. Karthikeyan, and K. Jayamoorthy (2016). Effect of bulk and nano-Fe2O3 particles on peanut plant leaves studied by Fourier transform infrared spectral studies. J. Adv. Res. 7(5): 739-747.
- Thompson, J.F. (1980). 10 - Arginine synthesis, proline synthesis, and related processes. In: Amino Acids and Derivatives. Editor(s): B.J. Miflin, Academic Press. pp: 375-402.
- Uma Maheswari Devi, P., K. Vijaya Lakshmi, K. Janardhan Reddy, and P. Reddanna (2005). Differential expression of lipoxygenase in green gram (Vigna radiata). Legume Res. 28(2): 87-93.
- Vaughn, M.W., M. Tanurdžić, Z. Lippman, H. Jiang, R. Carrasquillo, P.D. Rabinowicz, N. Dedhia, W.R. McCombie, N. Agier, Bulski, V. Colot, R.W. Doerge, and R.A. Martienssen (2007). Epigenetic natural variation in Arabidopsis thaliana. PLoS Biol. 5(7), e174: 1617-1629.
- Wang, W.H., B. Kohler, F.Q. Cao, and L.H. Liu (2008). Molecular and physiological aspects of urea transport in higher plants. Plant Sci. 175(4): 467-477.
- Xiong, L.Z., C.G. Xu, M.A. S. Maroof, and Q.F. Zhang (1999). Patterns of cytosine
methylation in an elite rice hybrid and its parental lines, detected by a
methylation-sensitive amplification polymorphism technique. Mol. Gen. Genet. 261(3): 439-446.
- Zluvova, J., J. Bohuslav, and B. Vyskot (2001). Immunohistochemical study of DNA methylation during plant development. J. Exp. Bot. 52(365): 2265-2273.
- Zonia, L.E., N.E. Stebbins, and J.C. Polacco (1995). Essential role of urease in germination of nitrogen-limited Arabidopsis thaliana Plant Physiol. 107(4): 1097-1103.
|





